Code
library(tidyverse)
library(haven)
library(fixest)
library(didimputation)
library(broom)
library(ggplot2)
# Chargement des données
d97 <- read_rds("data/derived/data_matched_1997.rds") %>%
rename(spei_wc_n_2 = spei_wc_1995,
spei_wc_n_1 = spei_wc_1996,
spei_wc_n = spei_wc_1997) %>%
mutate(hv219 = zap_labels(hv219), # hhh sex (1/2)
hv220 = zap_labels(hv220)) # hhh age (num)
d08 <- read_rds("data/derived/data_matched_2008.rds") %>%
rename(spei_wc_n_2 = spei_wc_2006,
spei_wc_n_1 = spei_wc_2007,
spei_wc_n = spei_wc_2008) %>%
mutate(hv219 = zap_labels(hv219),
hv220 = zap_labels(hv220))
d11 <- read_rds("data/derived/data_matched_2011.rds") %>%
rename(spei_wc_n_2 = spei_wc_2009,
spei_wc_n_1 = spei_wc_2010,
spei_wc_n = spei_wc_2011) %>%
mutate(hv219 = zap_labels(hv219),
hv220 = zap_labels(hv220))
d13 <- read_rds("data/derived/data_matched_2013.rds") %>%
rename(spei_wc_n_2 = spei_wc_2011,
spei_wc_n_1 = spei_wc_2012,
spei_wc_n = spei_wc_2013) %>%
mutate(hv219 = zap_labels(hv219),
hv220 = zap_labels(hv220))
d16 <- read_rds("data/derived/data_matched_2016.rds") %>%
rename(spei_wc_n_2 = spei_wc_2014,
spei_wc_n_1 = spei_wc_2015,
spei_wc_n = spei_wc_2016) %>%
mutate(hv219 = zap_labels(hv219),
hv220 = zap_labels(hv220))
d21 <- read_rds("data/derived/data_matched_2021.rds") %>%
rename(spei_wc_n_2 = spei_wc_2019,
spei_wc_n_1 = spei_wc_2020,
spei_wc_n = spei_wc_2021) %>%
mutate(hv219 = zap_labels(hv219),
hv220 = zap_labels(hv220))
# Préparation des données
dat <- bind_rows(d97, d08, d11, d13, d16, d21) %>%
filter(GROUP %in% c("Treatment","Control")) %>%
mutate(
hv219 = factor(hv219, levels = c(1,2), labels = c("Homme","Femme")), # sexe (cat.)
hv220 = as.numeric(hv220), # âge
treat = as.integer(GROUP == "Treatment"),
w_svy = hv005 / 1e6,
w_all = w_svy * weights, # poids d'enquête × poids de matching (si 'weights' existe)
id = row_number(),
# Map des années de statut -> première année d'observation post (treatment_phase)
treatment_phase = case_when(
STATUS_YR == 2010 ~ 2011,
STATUS_YR == 2012 ~ 2013,
STATUS_YR == 2015 ~ 2016,
STATUS_YR == 2017 ~ 2021,
is.na(STATUS_YR) ~ 0,
TRUE ~ STATUS_YR
)
)
# Outcome h
yvar <- "wealth_centile_rural_weighted"
# fixest DID 2×2: placebo 1997–2008, traitement 2008–2021 ------------------
# Placebo 1997–2008
pre <- dat %>%
filter(DHSYEAR %in% c(1997, 2008)) %>%
mutate(post = as.integer(DHSYEAR == 2008),
treat_post = treat * post)
f_pre <- as.formula(paste(
yvar, "~ treat + post + treat_post +",
"spei_wc_n_2 + spei_wc_n_1 + spei_wc_n + hv219 + hv220"
))
m_pre <- feols(f_pre, data = pre, weights = ~ w_all, cluster = ~ hv001)
# Traitement 2008–2021
main <- dat %>%
filter(DHSYEAR %in% c(2008, 2021)) %>%
mutate(post = as.integer(DHSYEAR == 2021),
treat_post = treat * post)
f_main <- f_pre # même formule
m_main <- feols(f_main, data = main, weights = ~ w_all, cluster = ~ hv001)
etable(m_pre, m_main, headers = c("Placebo 97–08", "Traitement 08–21")) m_pre m_main
Placebo 97–08 Traitement 08–21
Dependent Var.: wealth_centile_rural_weighted wealth_centile_rural_weighted
Constant 44.96*** (4.313) 37.89*** (2.715)
treat -1.797 (4.156) 1.484 (3.454)
post -6.394 (6.043) 22.55*** (4.543)
treat_post 3.736 (5.392) -2.751 (4.300)
spei_wc_n_2 2.114 (3.907) -5.482. (3.310)
spei_wc_n_1 -1.055 (2.639) -8.214*** (2.203)
spei_wc_n 2.411 (4.317) 17.72*** (2.512)
hv219Femme -1.735. (0.9367) -3.268*** (0.7580)
hv220 0.0966*** (0.0255) 0.0921*** (0.0182)
_______________ _____________________________ _____________________________
S.E.: Clustered by: hv001 by: hv001
Observations 6,831 11,686
R2 0.00655 0.08010
Adj. R2 0.00539 0.07948
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Code
# Extraction compacte des deux effets DID
did_row <- function(model, year_post, vc = ~ hv001, term = "treat_post"){
summary(model, vcov = vc) %>% broom::tidy() %>%
filter(term == !!term) %>%
transmute(year = year_post, estimate, se = std.error)
}
did_df <- bind_rows(
did_row(m_pre, 2008),
did_row(m_main, 2021)
) %>%
mutate(period = factor(ifelse(year == 2008, "1997–2008", "2008–2021"),
levels = c("1997–2008", "2008–2021")),
lo = estimate - 1.96*se,
hi = estimate + 1.96*se)
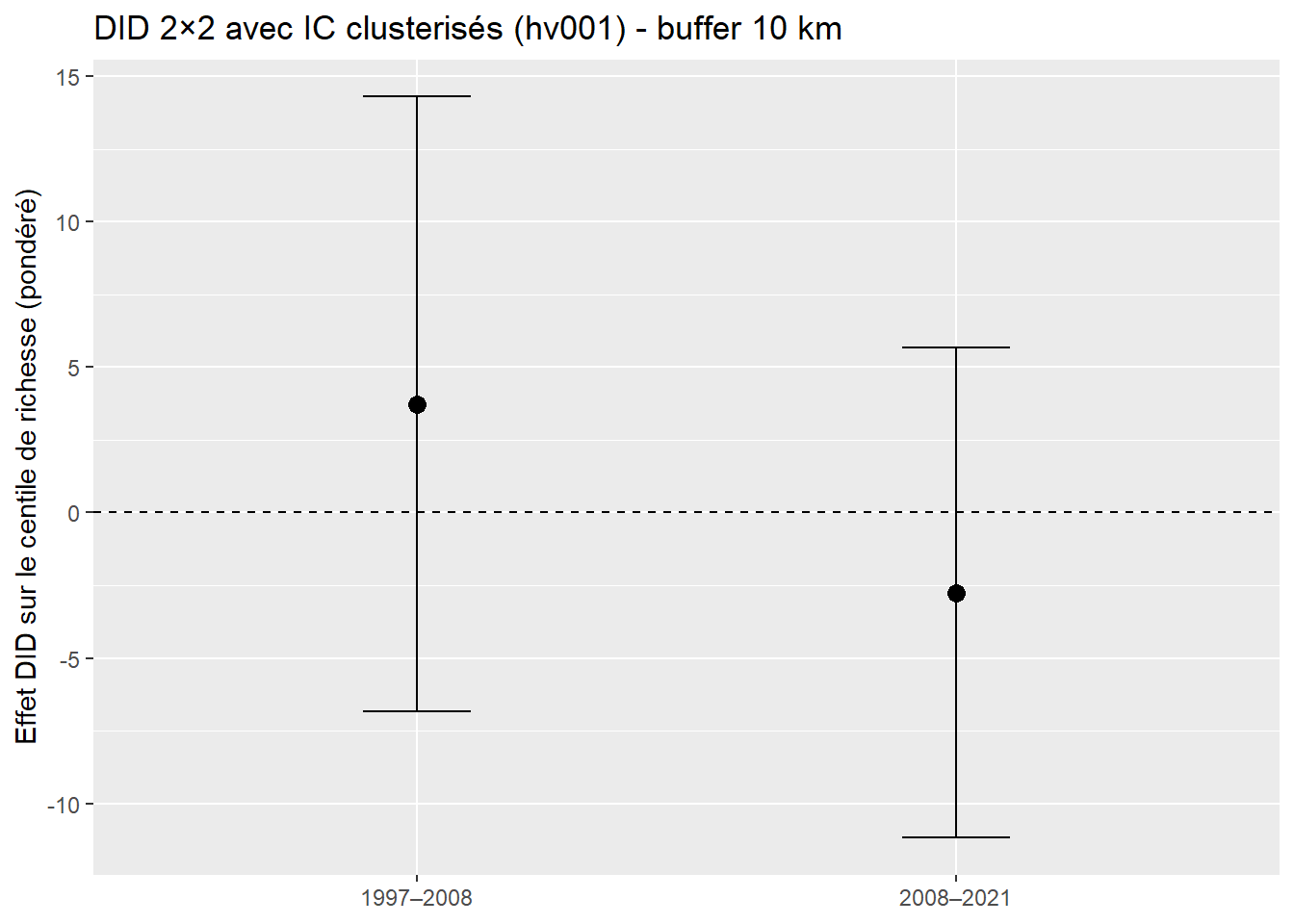
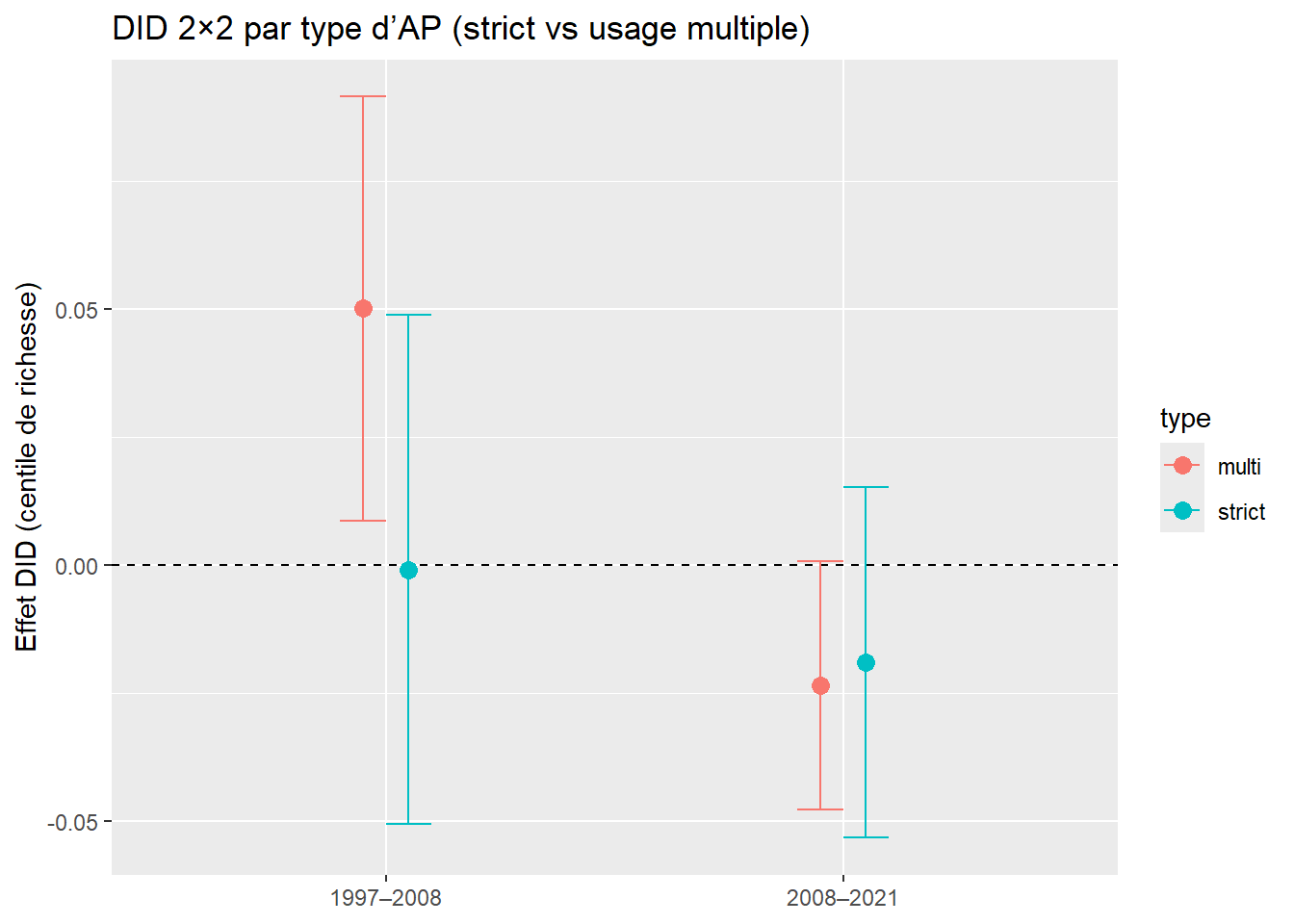
ggplot(did_df, aes(x = period, y = estimate)) +
geom_hline(yintercept = 0, linetype = "dashed") +
geom_errorbar(aes(ymin = lo, ymax = hi), width = .2) +
geom_point(size = 3) +
labs(x = NULL, y = "Effet DID sur le centile de richesse (pondéré)",
title = "DID 2×2 avec IC clusterisés (hv001) - buffer 10 km")